Logistic Regression (Part III)
University of Cincinnati
Variations of logistic regression
Logistic regression
In logistic regression, we use the logit: \[ \text{logit}\left(p\right) = \boldsymbol{x}^\top\boldsymbol{\beta} = \eta \] which results in \[ p = \left[1 + \exp\left(-\eta\right)\right]^{-1} = F\left(\eta\right) \]
Technically, \(F\) can be any monotonic function that maps \(\eta\) to \(\left[0, 1\right]\) (e.g., any CDF will work)
\(F^{-1}\) is called the link function
The probit model
The probit model uses \(F\left(\eta\right) = \Phi\left(\eta\right)\) (i.e., the CDF of a standard normal distribution) \[ P\left(Y = 1 | \boldsymbol{x}\right) = \Phi\left(\beta_0 + \beta_1x_1 + \cdots\right) = \Phi\left(\boldsymbol{x}^\top\boldsymbol{\beta}\right) \]
\(F^{-1} = \Phi^{-1}\) is called the probit link, which yields \[ \text{probit}\left(p\right) = \Phi^{-1}\left(p\right) = \boldsymbol{x}^\top\boldsymbol{\beta} \]
The term probit is short for probability unit
Proposed in Bliss (1934) and still common for modeling dose-response relationships
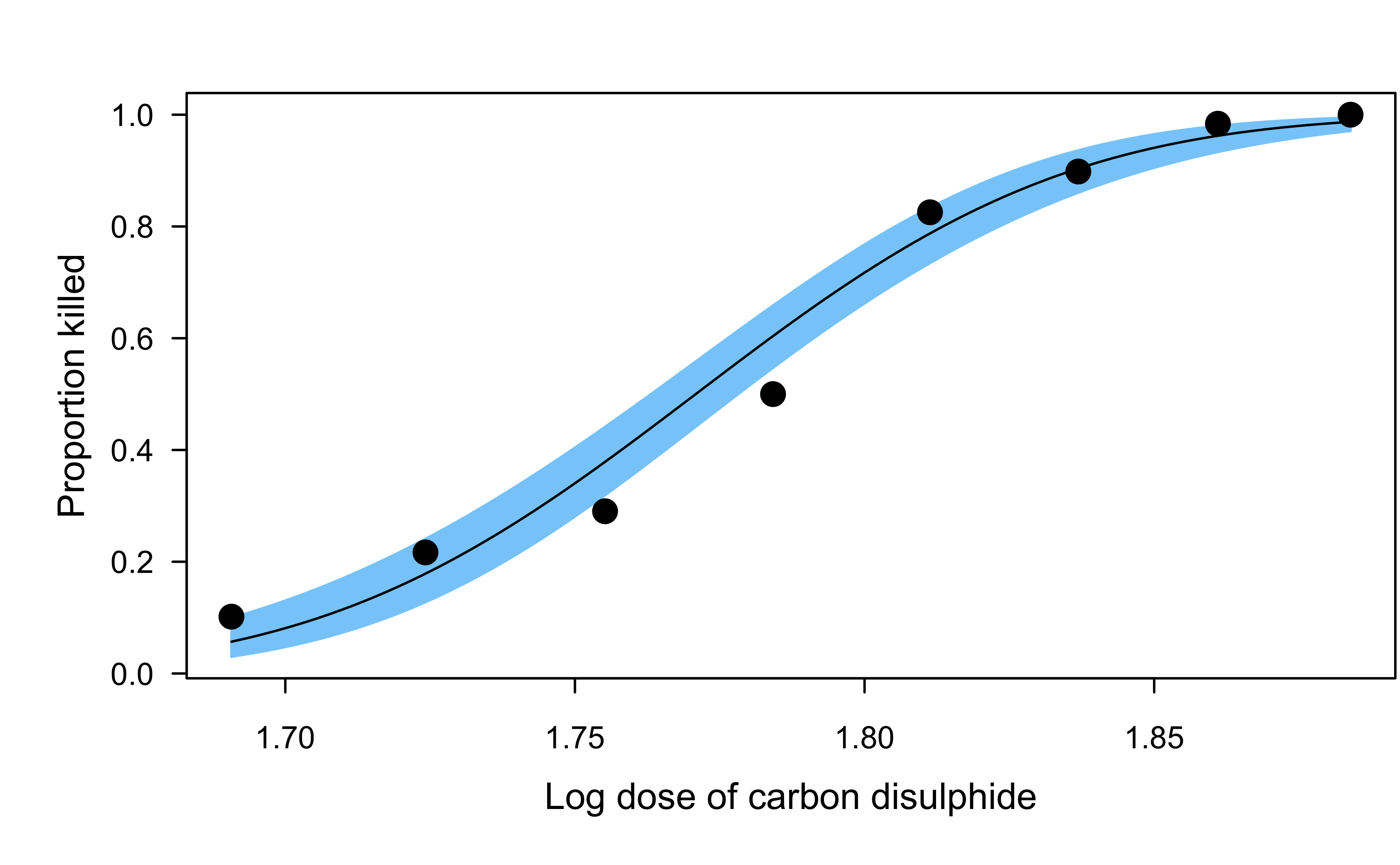
Dobson’s beetle data
The data give the number of flour beetles killed after five hour exposure to the insecticide carbon disulphide at eight different concentrations.
Other link functions
Common link function for binary regression include:
- Logit (most common and the default an most software)
- Probit (next most common)
- Log-log
- Complimentary log-log
- Cauchit
Application to wcgs data set
Comparing coefficients from different link functions:
Show R code
wcgs <- na.omit(subset(faraway::wcgs, select = -c(typechd, timechd, behave)))
# Fit binary GLMs with different link functions
fit.logit <- glm(chd ~ ., data = wcgs, family = binomial(link = "logit"))
fit.probit <- glm(chd ~ ., data = wcgs, family = binomial(link = "probit"))
fit.cloglog <- glm(chd ~ ., data = wcgs, family = binomial(link = "cloglog"))
fit.cauchit <- glm(chd ~ ., data = wcgs, family = binomial(link = "cauchit"))
# Compare coefficients
coefs <- cbind(
"logit" = coef(fit.logit),
"probit" = coef(fit.probit),
"cloglog" = coef(fit.cloglog),
"cauchit" = coef(fit.cauchit)
)
round(coefs, digits = 3) logit probit cloglog cauchit
(Intercept) -12.246 -6.452 -11.377 -24.086
age 0.062 0.031 0.057 0.115
height 0.007 0.003 0.007 0.053
weight 0.009 0.005 0.007 0.003
sdp 0.018 0.010 0.017 0.042
dbp -0.001 -0.001 -0.001 -0.001
chol 0.011 0.006 0.010 0.020
cigs 0.021 0.011 0.019 0.043
dibepB -0.658 -0.341 -0.604 -1.320
arcuspresent 0.211 0.101 0.206 0.711Application to wcgs data set
Comparing fitted values from different link functions:
Show R code
logit probit cloglog cauchit
1 0.071 0.072 0.072 0.076
2 0.073 0.074 0.075 0.084
3 0.010 0.006 0.011 0.038
4 0.010 0.006 0.012 0.039
5 0.169 0.167 0.168 0.150
6 0.034 0.033 0.035 0.051Application to wcgs data set
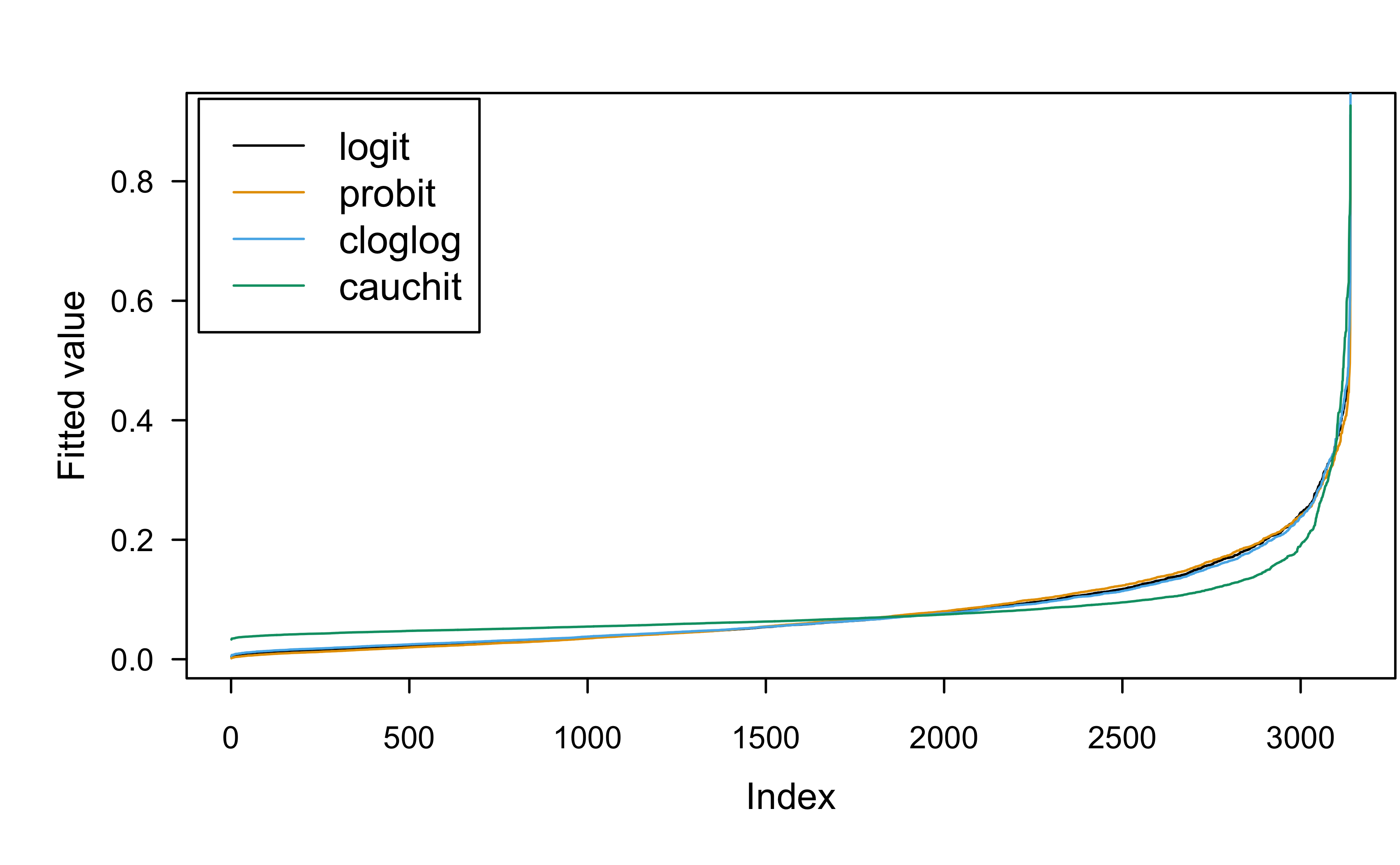
Comparing fitted values from different link functions:
Show R code
As a latent variable model
Logistic regression (and the other link functions) has an equivalent formulation as a latent variable model
Consider a linear model with continuous outcome \(Y^\star = \boldsymbol{x}^\top\boldsymbol{\beta} + \epsilon\)
Imagine that we can only observe the binary variable \[ Y = \begin{cases} 1 \quad \text{if } Y^\star > 0, \\ 0 \quad \text{otherwise} \end{cases} \]
Assuming \(\epsilon \sim \text{Logistic}\left(0, 1\right)\) leads to the usual logit model
Assuming \(\epsilon \sim N\left(0, 1\right)\) leads to the probit model
And so on…
Application: surrogate residual
- There are several types of residuals defined for logistic regression (e.g., deviance residuals and Pearson residuals)
- The surrogate residual act like the usual residual from a normal linear model and has similar properties!
- See Liu and Zhang (2018), Greenwell et al. (2018), and Cheng et al. (2020) for details
- A novel R-squared measure based on the surrogate idea was proposed in Liu et al. (2022)
- For implementation, see R packages sure, SurrogateRsq, and PAsso
Binomial data
O-rings example
On January 28, 1986, the Space Shuttle Challenger broke apart 73 seconds into its flight, killing all seven crew members aboard. The crash was linked to the failure of O-ring seals in the rocket engines. Data was collected on the 23 previous shuttle missions. The launch temperature on the day of the crash was 31 \(^\circ\)F.
O-rings example
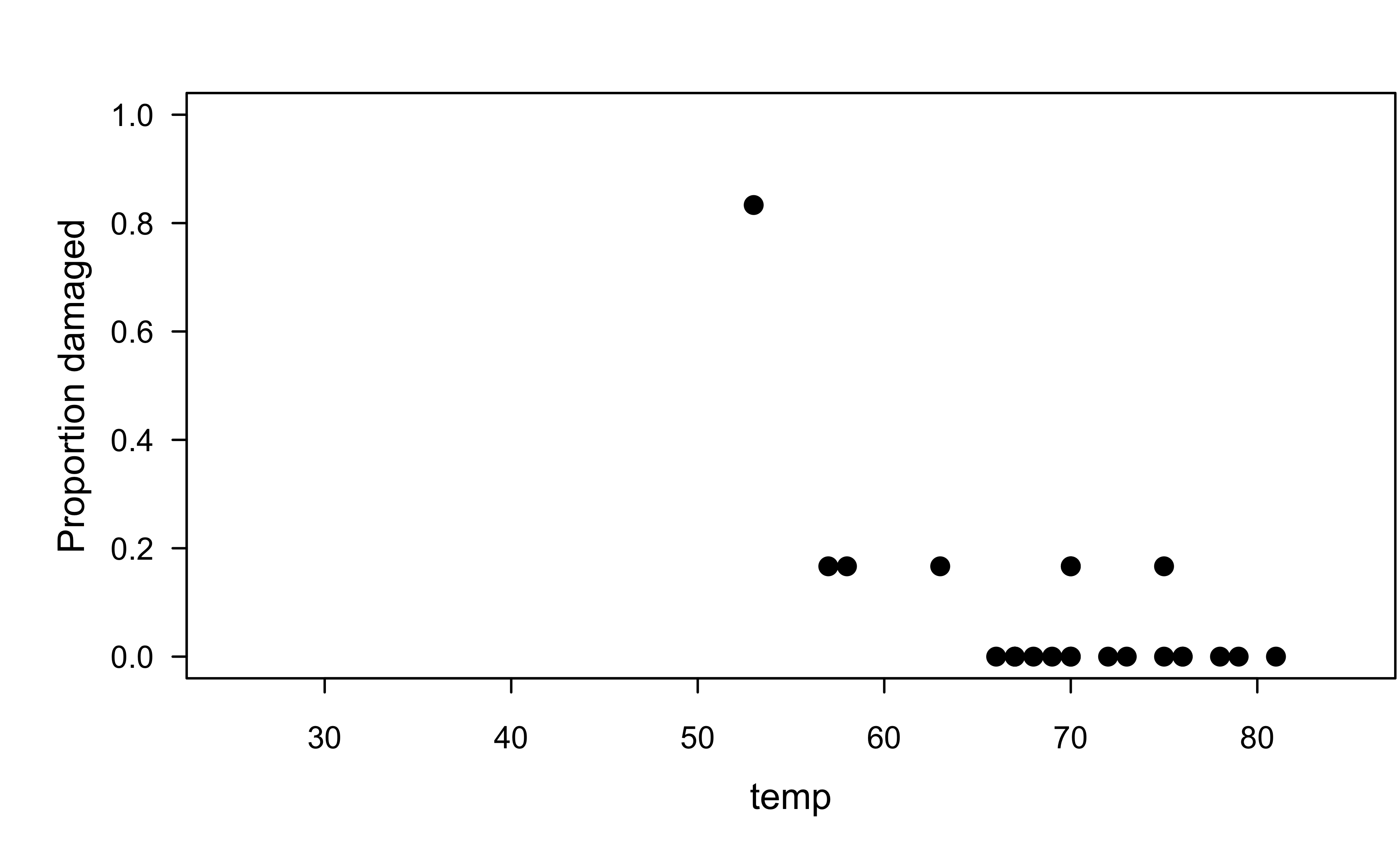
O-rings example
Expand binomial data into independent Bernoulli trials
O-rings example
Here we’ll just fit a logistic regression to the expanded bernoulli version of the data
Show R code
Call:
glm(formula = damage ~ temp, family = binomial(link = "logit"),
data = orings2)
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 11.66299 3.29616 3.538 0.000403 ***
temp -0.21623 0.05318 -4.066 4.77e-05 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 76.745 on 137 degrees of freedom
Residual deviance: 54.759 on 136 degrees of freedom
AIC: 58.759
Number of Fisher Scoring iterations: 6O-rings example
What’s the estimated probability that an O-ring will be damaged at 31 \(^\circ\)F? Give a point estimate as well as a 95% confidence interval.
$fit
1
0.9930342
$se.fit
1
0.01153302
$residual.scale
[1] 1Show R code
[1] 0.8444824 0.9997329O-rings example
Show R code
# Is this extrapolating?
plot(damage / 6 ~ temp, data = orings, pch = 19, cex = 1.3,
col = adjustcolor(1, alpha.f = 0.3), xlim = c(0, 100), ylim = c(0, 1))
x <- seq(from = 0, to = 100, length = 1000)
y <- predict(orings.lr, newdata = data.frame("temp" = x),
type = "response")
lines(x, y, lwd = 2, col = 2)
abline(v = 31, lty = 2, col = 3)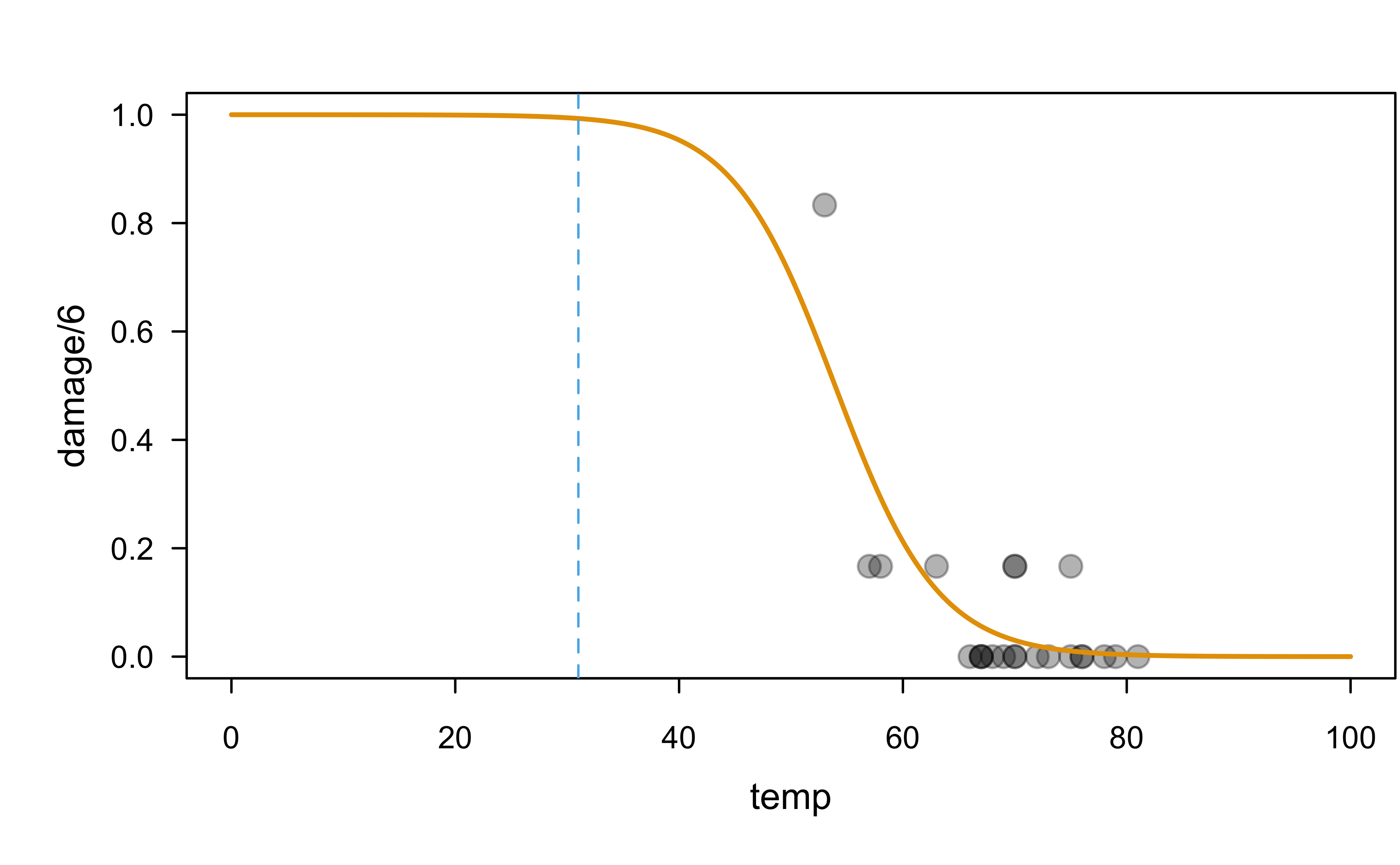
O-rings examples
More interesting question: at what temperature(s) can we expect the risk/probability of damage to exceed 0.8?
This is a problem of inverse estimation, which is the purpose of the investr package!
O-rings example
Here’s an equivalent logistic regression model fit to the original binomial version of the data (need to provide number of successes and number of failures)
Show R code
Call:
glm(formula = cbind(damage, 6 - damage) ~ temp, family = binomial(link = "logit"),
data = orings)
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 11.66299 3.29626 3.538 0.000403 ***
temp -0.21623 0.05318 -4.066 4.78e-05 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Dispersion parameter for binomial family taken to be 1)
Null deviance: 38.898 on 22 degrees of freedom
Residual deviance: 16.912 on 21 degrees of freedom
AIC: 33.675
Number of Fisher Scoring iterations: 6Overdispersion
Overdispersion
For a Bernoulli random variable \(Y\), \(E(Y) = p\) and \(V(Y) = p(1 - p)\).
Sometimes the data exhibit variance greater than expected. Adding a dispersion parameter makes the model more flexible: \(V(Y) = \sigma^2 p(1 - p)\).
Overdispersion occurs when variability exceeds expectations under the response distribution. Underdispersion is less common.
Overdispersion
For a correctly specified model, the Pearson chi-square statistic and deviance, divided by their degrees of freedom, should be about one. Values much larger indicate overdispersion.
Such goodness-of-fit statistics require replicated data. Problems like outliers, wrong link function, omitted terms, or untransformed predictors can inflate goodness-of-fit statistics.
A large difference between the Pearson statistic and deviance suggests data sparsity.
O-rings example
You can check for overdispersion manually or by using performance:
Overdispersion
Two common (but equivalent) ways to handle overdispersion in R:
Estimate the dispersion parameter \(\sigma^2\) and provide it to
summary()to adjust the standard errors approriatelyUse the
quasibinomial()family
Overdispersion
Estimate the dispersion parameter \(\sigma^2\); analagous to MSE in linear regression
[1] 1.336542Show R code
Call:
glm(formula = cbind(damage, 6 - damage) ~ temp, family = binomial(link = "logit"),
data = orings)
Coefficients:
Estimate Std. Error z value Pr(>|z|)
(Intercept) 11.66299 3.81077 3.061 0.002209 **
temp -0.21623 0.06148 -3.517 0.000436 ***
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Dispersion parameter for binomial family taken to be 1.336542)
Null deviance: 38.898 on 22 degrees of freedom
Residual deviance: 16.912 on 21 degrees of freedom
AIC: 33.675
Number of Fisher Scoring iterations: 6Overdispersion
Can use quasibinomial() family to account for over-dispersion automatically (notice the estimated coeficients don’t change)
Show R code
Call:
glm(formula = cbind(damage, 6 - damage) ~ temp, family = quasibinomial,
data = orings)
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 11.66299 3.81077 3.061 0.00594 **
temp -0.21623 0.06148 -3.517 0.00205 **
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
(Dispersion parameter for quasibinomial family taken to be 1.336542)
Null deviance: 38.898 on 22 degrees of freedom
Residual deviance: 16.912 on 21 degrees of freedom
AIC: NA
Number of Fisher Scoring iterations: 6Generalized additive models (GAMs)
\(\text{logit}\left(p\right) = \beta_0 + f_1\left(x_1\right) + f_2\left(x_2\right) + \dots + f_p\left(x_p\right)\)
The \(f_i\) are referred to as shape functions or term contributions and are often modeled using splines
Can also include pairwise interactions of the form \(f_{ij}\left(x_i, x_j\right)\)
Easy to interpret (e.g., just plot the individual shape functions)!
- Interaction effects can be understood with heat maps, etc.
Modern GAMs, called GA2Ms, automatically include relevant pairwise interaction effects
Explainable boosting machines (EBMs)
- EBM ≈ .darkblue[GA2M] + .green[Boosting] + .tomato[Bagging]
- Python library: interpret
Questions?

BANA 7042: Statistical Modeling